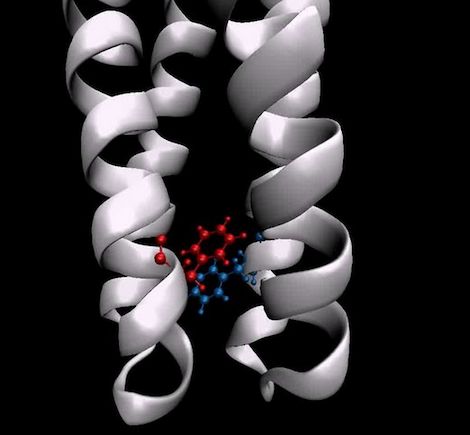
Over at OLCF, Katie Elyce Jones writes that researchers are using the GPU-powered Titan supercomputer to study a key molecular switch that controls cell behavior. While scientists want to manipulate cell behavior for many reasons, this ability would enable them to cripple cells and pathogens that are causing disease in the body.
Researchers discovered the molecular switch by simulating 140,000 atoms that make up the signaling part of the Tsr chemoreceptor dimer that controls motility in E. coli. A dimer is a system of two identical receptor molecules. Like other receptors, Tsr spans the cell membrane, communicating to proteins inside the cell in order to respond to threats or opportunities in the environment. Exactly how receptors send signals to neighboring proteins is a critical, unanswered question in molecular cell biology. The results, published in Nature Communications, directly address this question and stand apart from previous research because of the computational power applied to the problem.
While the team is currently running a trimer of dimers simulation on Titan, it is also exploring how much computational power it will take to simulate even larger receptor complexes than a single trimer. While these kinds of simulations lie farther in the future, Ortega said, systems like Titan are making planning for them possible.
Read the Full Story.