Although the underlying hardware architecture is critical to efficiency, the management of the data flow becomes paramount to success. SGI’s Data Management Framework (DMF) software – when used within personalized medicine applications – provides a large-scale, storage virtualization and tiered data management platform specifically engineered to administer the billions of files and petabytes of structured and unstructured fixed content generated by highly scalable and extremely dynamic life sciences applications.
When used in conjunction with the SGI UV300 converged compute and flash-based storage environments shown in the optimized high-performance workflow model (Figure 4), DMF is able to automatically manage data within the system in order to meet the needs of the executing workflows.
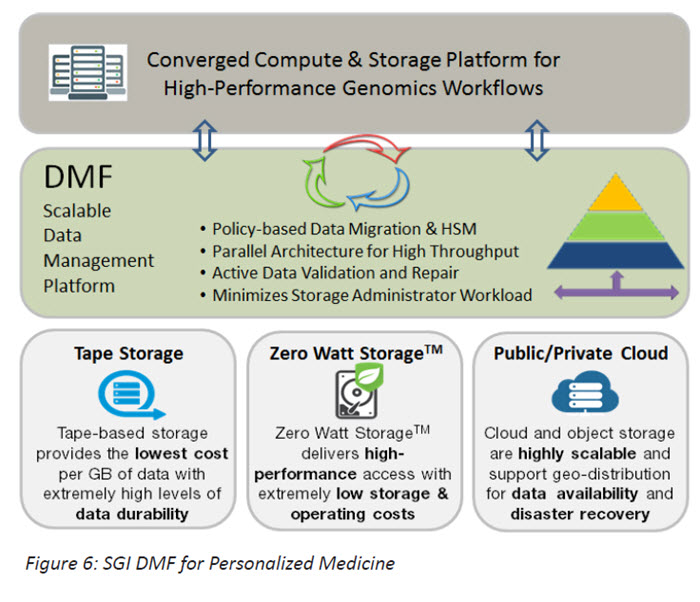
The DMF architecture consists of interface, policy management, and storage management layers that administer a variety of media tiers. The interface layer for applications is a standard POSIX file system that can be leveraged within existing compute and storage environments with no requirement for researchers to modify their applications. DMF transparently integrates within the file system environment and uses storage management policies that can be tuned for each research and lab environment to silently and automatically move data to alternative tiers of storage that can include cloud, object storage, disk-based cold storage and even library-based tape.
Major research institutes and laboratories trust DMF to manage, backup, archive, and protect their critical biomedical data. As a policy-driven, centralized data management platform, DMF provides:
- Self-Healing for Long Term Data Assurance: High data availability and accessibility to vital life sciences information, including genomic sequencing, drug discovery, drug therapy, neurobiology, and clinical research data.
- Petascale Architecture: Archive scalability to multiple petabytes with no performance degradation to accommodate the ever-increasing volume of biomedical information.
- Broad Array of Storage Options: Cost-effective operation and reduced total cost of data ownership by storing information on different media types according to access requirements and business value.
- No Separate Backup Requirements: Enhanced business continuance through integrated backups, data replication, and multiple disaster recovery strategies that replace system vulnerability with strengthened researcher confidence and data protection.
- Integration with Existing Applications and Tools: Data sharing by diverse research units across the organization to ensure unified access to the same set of critical information.
Powering the Next Phase of Research and Results
In just a few short years genomic sequencing, including DNA sequencing and whole genome analysis have had a profound impact on bioinformatics in plant, animal and microbial science. It has enabled significant advances in research disciplines including the environment, food, and healthcare. At the same time the amount of data to be captured, stored, analyzed and interpreted is growing exponentially. Advances in next-generation sequencing will greatly lower costs and increase productivity, allowing organizations of all sizes to perform many experiments in a month. This requires the management of workflows with terabytes of data for a single sequencing run and petabytes for an entire laboratory.
To read 5 case studies on how genomics is enabling personalized medicine download the insideHPC Guide to Personalized Medicine and Genomics, courtesy of SGI and Intel.




