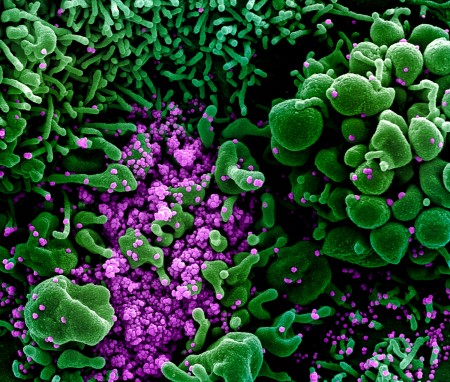
This NIH image, unrelated to NERSC’s participation in COVID-19 research, was captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Select image to enlarge. (Credit: National Institute of Allergy and Infectious Diseases, NIH)
As public health officials respond in real-time to the global pandemic of COVID-19 and countries around the world increase restrictions and issue shelter-in-place orders to stem the rising tide of this disease, cases continue to increase, with more than 1 million reported and 52,000 deaths as of April 2, 2020. Scientists are working around the clock to understand this respiratory virus, which exhibits characteristics similar to several viral epidemics such as the acute respiratory syndrome coronavirus (SARS-CoV) epidemic in 2002 to 2003 and the Middle East respiratory syndrome coronavirus (MERS-CoV) that began in Saudi Arabia in 2012.
In early January, the Chinese scientists who sequenced the viral COVID-19 genome made that data public, and now researchers are scrutinizing this data for insight into the disease by using supercomputing resources to meet the challenge.
For example, City of Hope’s Beckman Research Institute, which is already involved with a number of COVID-19 genome analyses and related projects and City of Hope affiliate TGen (Translational Genomics Research Institute) is using supercomputing resources at Lawrence Berkeley National Laboratory’s (Berkeley Lab) National Energy Research Scientific Computing Center (NERSC) to aid in these efforts.
The collaboration with City of Hope will use the computing resources of NERSC to identify small molecules that can be used to assist in the diagnosis of COVID-19, design peptide libraries to mimic virus-host interactions, and annotate the virulence of the Italian strain as compared to the Chinese strain of COVID-19,” said Blake Simmons, division director of Biological Systems and Engineering at Berkeley Lab.
In addition to the collaboration with the City of Hope, NERSC is a member of the COVID-19 High Performance Computing Consortium. In support of the Consortium, NERSC has reserved a portion of its Director’s Discretionary Reserve time on Cori, a Cray XC40 supercomputer, to support COVID-19 research efforts. The GPU partition on Cori was installed to help prepare applications for the arrival of Perlmutter, NERSC’s next-generation system that is scheduled to begin arriving later this year and will rely on GPUs for much of its computational power.
NERSC is pleased to be able to support such vital research at this time of critical need,” said Richard Gerber, Senior Science Advisor for NERSC, a Department of Energy Office of Science user facility. “Our staff has already set up procedures to quickly enable access to our computational and data resources and is on call to provide expert guidance on how to best use them.”
The City of Hope research targets the entire viral genome, with particular scrutiny on key viral proteins, initially focused on the Spike and M protein, as potential drug targets with the hope of developing antiviral treatments.
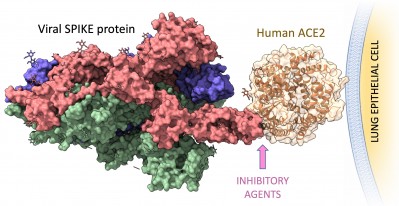
Crystal structure of SARS-Cov-2 SPIKE protein in complex with human ACE2 on the surface of the lung epithelial cells. The binding interface is the target for the computational design of inhibitory agents to prevent viral infection. Select image to enlarge. (Credit: Supriyo Bhattacharya)
Designing inhibitory agents, such as peptides targeting viral proteins, requires performing long-timescale molecular dynamics simulations on multiple proteins, some of which are huge and complex,” said Supriyo Bhattacharya, assistant professor at Translational Bioinformatics division of the Center for Informatics, a part of Beckman Research Institute at City of Hope. “To achieve this, the simulations needed support from GPU-accelerated computing, which NERSC is able to offer us. We will run these simulations on NERSC’s GPU resources and use the simulation data to design peptides which will be tested by our experimentalist collaborators in high-throughput assays.”
For this project, the researchers are employing the GROMACS and Amber codes. GROMACS is a versatile package created to perform molecular dynamics and is well-suited to this research as it was designed for biochemical molecules like proteins, lipids, and nucleic acids that have a lot of complicated bonded interactions. As well, GROMACS is extremely fast at calculating the non-bonded interactions that usually dominate simulations. The Amber suite of biomolecular simulation tools is a set of molecular mechanical force fields for the simulation of biomolecules and it is also a package of molecular simulation programs, including source code and demos.
For more than 45 years, scientists have tapped into NERSC’s systems and expertise to tackle some of the most challenging problems, but this is the first time that we have dedicated resources to fighting a medical emergency that is threatening the globe,” said Sudip Dosanjh, director of NERSC. “This is truly new territory for us, and we’re doing all we can to support this effort.”
Source: Carol Pott at NERSC