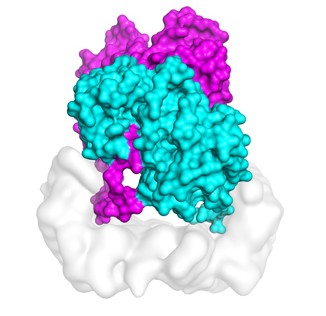
Molecular dynamics simulations on TACC’s Stampede2 supercomputer tested the stability of the structure of B-Raf:14-3-3 complex, which when mutated is linked to skin cancer.
In this TACC podcast, UC Berkeley scientists describe how they are using powerful supercomputers to uncover the mechanism that activates cell mutations found in about 50 percent of melanomas. They’re hopeful their study can help lead to a better understanding of skin cancer and to the design of better drugs.
Deepti Karandur, postdoctoral researcher, John Kuriyan Lab at UC Berkeley; postdoctoral fellow at the Howard Hughes Medical Institute.In 2002, scientists found a link between skin cancer and mutations of B-Raf (Rapidly Accelerated Fibrosarcoma) kinase, a protein that’s part of the signal chain that starts outside the cell and goes inside to direct cell growth. This signal pathway, called the Ras/Raf/Mek/Erk kinase pathway, is important for cancer research, which seeks to understand out of control cell growth. According to the study, about 50 percent of melanomas have a specific single mutation on B-Raf, known as the valine 600 residue to glutamate (V600E).
B-Raf V600E thus became an important drug target, and specific inhibitors of the mutant were developed in the following years. The drugs inhibited the mutant, but something strange happened. Paradoxically, quieting the mutant had a down side. It activated the un-mutated, wild-type B-Raf protein kinases, which again triggered melanoma.
With this background, we worked on studying the structure of this important protein, B-Raf,” said Yasushi Kondo, a postdoctoral researcher in the John Kuriyan Lab at UC Berkeley. Kondo is the co-author of an October 2019 study in the journal Science that determined the structure of the complex of proteins that make up B-Raf and also found how the paradoxical B-Raf activation happens. “We aimed to study the more native-like state of the protein to understand how it’s regulated in the cells, because most of the studies have been focused on the isolated kinase domain and how the drugs bind to the kinase domain.”
Kondo’s and his colleagues used cryo-electron microscopy (cryo-EM) to determine the structure of B-Raf 14-3-3 complex, basically cryogenically freezing the protein complex, which kept it in a chemically-active, near-natural environment. Next they flashed it with electron beams to obtain thousands of ‘freeze frames.’ They sifted out background noise and reconstructed three-dimensional density maps that showed previously unknown details in the shape of the molecule. And for proteins, form follows function.
Kondo explained that the structure revealed an asymmetric organization of the complex, formed by two sets of internally symmetrical dimers, or pairs of bonded molecules. “We propose that this unexpected arrangement enables asymmetric activation of the B-Raf dimer, which is a mechanism that provides an explanation of the origin of the paradoxical activation of B-Raf by small molecule inhibitors,” Kondo said.
The study’s computational challenges involved molecular dynamics simulations that modeled the protein at the atomic level, determining the forces of every atom on every other atom for a system of about 200,000 atoms at time steps of two femtoseconds.
For small systems, we can see what’s happening relatively quickly, but for large systems like these, especially large biomolecular systems, these changes happen on like nanosecond timescales, microsecond timescales, or even millisecond timescales,” Karandur said.

The Stampede2 supercomputer at the Texas Advanced Computing Center is an allocated resource of the Extreme Science and Engineering Discovery Environment (XSEDE) funded by the National Science Foundation (NSF).
The Stampede2 supercomputer at the Texas Advanced Computing Center is an allocated resource of the Extreme Science and Engineering Discovery Environment (XSEDE) funded by the National Science Foundation (NSF).Karandur and colleagues turned to XSEDE, the NSF-funded Extreme Science and Engineering Discovery Environment, for allocation time on the Stampede2 supercomputer at the Texas Advanced Computing Center (TACC) to do the simulations, as well as the Bridges system at the Pittsburgh Supercomputing Center to investigate other proteins in the pathway. Stampede2’s Skylake processor nodes, networked with Intel Omnipath, made quick work of the optimized-for-supercomputers NAMD molecular dynamics simulations.
Stampede2 runs very, very fast, and it’s very efficient. We generated a total of about 1.5 microseconds of trajectories for our systems in about four to six weeks. Whereas, if we ran it on our own in-house cluster it would have taken us months or longer,” Karandur said.
About XSEDE, Karandur commented: “I think it’s an amazing resource. I’ve been running simulations starting from when I was a graduate student. XSEDE made it possible for us to access timescales that are biologically relevant. Everything that happens in a cell, happens on microsecond timescales, to millisecond timescales, to longer. When I was starting, we could not run this simulation on any system anywhere. I mean, it would have taken five years, or more. To be able to do it in weeks and say, okay, we know understand why this is important so we can now start to gain real understanding into how the biology happens, is just amazing,” Karandur said.
And there remains a lot to be discovered about B-Raf. It’s just one link in the signal chain that governs cell growth and cancer.
“The structure that was resolved in this paper is part of a large, multi-domain system,” Karandur explained. “We don’t know what this complete protein looks like. We don’t see it in the structure. We don’t know what its dynamics look like, and how all these other parts of the protein play a role in maintaining the active state, or converting it from the inactive state to the active state.”
She furthered that as the system gets bigger, the pertinent structural changes happen over longer timescales, and bigger supercomputers are needed to handle the complexity, such as the NSF-funded Frontera supercomputer, also at TACC.
Frontera is getting there. We’re very excited about this. We are in the process of getting an allocation on Frontera,” Karandur said.
For non-scientists, this fundamental research could yield insight leading to better drugs for skin cancer.
The study, “Cryo-EM structure of a dimeric B-Raf:14-3-3 complex reveals asymmetry in the active sites of B-Raf kinases,” was published Oct. 4, 2019 in the journal Science. The authors are Yasushi Kondo, Deepti Karandur, John Kuriyan, and Kathryn Wong of UC Berkeley; Jana Ognjenović and Alan Merk of the National Cancer Institute; Saikat Banerjee, Kayla Kulhanek, Jeroen P. Roose of UC San Francisco; and Sriram Subramaniam of the University of British Columbia, Vancouver. Study funding came from the National Institutes of Health, the Aqueduct Foundation, the VGH & UBC Hospital Foundation, and the Government of Canada.
Stampede2 and Bridges are allocated resources of the Extreme Science and Engineering Discovery Environment (XSEDE) funded by the National Science Foundation (NSF).